

<- Back to Index
1.3 Pathways of phenylpropanoid biosynthesis
Go to:
|
|
Lignins are assembled from monolignols, which are synthesized via the shikimate
and phenylpropanoid pathways. Mutants in which the lignin biosynthetic genes
are affected have been identified in several species.
Hydroxycinnamic acids, which originate from the deamination, hydroxylation
and sometimes methylation of phenylalanine (and tyrosine in grasses), are
commonly incorporated into type II walls.
Click on Gene for links to the Wisconsin T-DNA project for development of homozygous insertional lines.
Click on hypertext links to download IR SPectra and Likely Spectrotypes.
COMT Group |
|||||||
| Gene | Annotation | Accession | Date
Received |
IR
Spectra |
PCs, % Classified | Likely Spectrotype | Comments |
| At5g54160 | COMT | SALK_135290 | 3/2/05 | wild(.xls) mutant(.xls) |
4 , 73 | ||
| At1g21100 | COMT-like1 | ||||||
| At1g21110 | COMT-like2 | ||||||
| At1g21120 | COMT-like3 | ||||||
| At1g21130 | COMT-like4 | ||||||
| At1g33030 | COMT-like5 | SALK_075264 | 3/2/05 | wild(.xls) mutant(.xls) |
4 , 86 | Yes | |
| At1g51990 | COMT-like6 | ||||||
| At1g63140 | COMT-like7 | ||||||
| At1g76790 | COMT-like8 | ||||||
| At1g77520 | COMT-like9 | ||||||
| At1g77530 | COMT-like10 | SALK_012948 | 3/2/05 | wild(.xls) mutant(.xls) |
4 , 80 | Yes | |
| At3g53140 | COMT-like11 | ||||||
| At5g37170 | COMT-like12 | SALK_059964 | 10/12/05 | wild(.xls) mutant(.xls) |
5 , 75 | ||
| At5g53810 | COMT-like13 | SALK_070607 | 3/2/05 | wild(.xls) mutant(.xls) |
4 , 76 | ||
| At4g35150 | O-methyltransferase Family 2 Protein | ||||||
| At4g35160 | O-methyltransferase Family 2 Protein | SALK_084490 | 3/2/05 | wild(.xls) mutant(.xls) |
4 , 87 | Yes | |
4CL Group |
|||||||
| Gene | Annotation | Accession | Date
Received |
IR
Spectra |
PCs, % Classified | Likely Spectrotype | Comments |
| At1g51680 | 4CL1 | ||||||
| At3g21240 | 4CL2 | ||||||
| At1g65060 | 4CL3 | SALK_003025 | 3/2/05 | wild(.xls) mutant(.xls) |
5 , 66 | ||
| At3g21230 | 4CL4 | ||||||
| At1g20510 | 4CL-like1 | ||||||
| At1g20500 | 4CL-like2 | SAIL_26_H04 | 3/1/05 | 4, 95 | |||
| At1g20490 | 4CL-like3 | ||||||
| At1g20480 | 4CL-like4 | SALK_001720 | 3/1/05 | 4, 86 | |||
| At1g62940 | 4CL-like5 | ||||||
| At4g19010 | 4CL-like6 | ||||||
| At4g05160 | 4CL-like7 | SALK_050214 | 3/2/05 | wild(.xls) mutant(.xls) |
4 , 83 | Yes | |
| At5g63380 | 4CL-like8 | SALK_130997 | 3/2/05 | wild(.xls) mutant(.xls) |
4 , 53 | ||
| At5g38120 | 4CL-like9 | ||||||
CCoAOMT Group |
|||||||
| Gene | Annotation | Accession | Date
Received |
IR
Spectra |
PCs, % Classified | Likely Spectrotype | Comments |
| At4g34050 | CCoAOMT1 | ||||||
| At1g24735 | CCoAOMT2 | ||||||
| At3g61990 | CCoAOMT3 | ||||||
| At3g62000 | CCoAOMT4 | ||||||
| At1g67990 | CCoAOMT5 | ||||||
| At1g67980 | CCoAOMT6 | ||||||
| At4g26220 | CCoAOMT7 | ||||||
PAL Group |
|||||||
| Gene | Annotation | Accession | Date
Received |
IR
Spectra |
PCs, % Classified | Likely Spectrotype | Comments |
| At2g37040 | PAL1 | SALK_022804 | 3/2/05 | wild(.xls) mutant(.xls) |
4 , 78 | ||
| At3g53260 | PAL2 | ||||||
| At5g04230 | PAL3 | ||||||
| At3g10340 | PAL4 | ||||||
CAD Group |
|||||||
| Gene | Annotation | Accession | Date
Received |
IR
Spectra |
PCs, % Classified | Likely Spectrotype | Comments |
| At4g39330 | CAD1 | SALK_037853 | 3/2/05 | wild(.xls) mutant(.xls) |
4 , 86 | Yes | |
| At3g19450 | CAD2 | ||||||
| At4g37970 | CAD3 | SALK_030496 | 3/2/05 | wild(.xls) mutant(.xls) |
5 , 87 | Yes | |
| At4g37980 | CAD4 | ||||||
| At4g37990 | CAD5 | ||||||
| At4g34230 | CAD6 | SALK_040062 | 10/12/05 | wild(.xls) mutant(.xls) |
5 , 71 | ||
| At2g21730 | CAD7 | ||||||
| At2g21890 | CAD8 | ||||||
| At1g72680 | CAD9 | ||||||
C3H & F5H & C4H Group |
|||||||
| Gene | Annotation | Accession | Date
Received |
IR
Spectra |
PCs, % Classified | Likely Spectrotype | Comments |
| At2g40890 | C3H1 | ||||||
| At1g74540 | C3H2 | ||||||
| At1g74550 | C3H3 | ||||||
| At2g30490 | C4H | ||||||
| At4g36220 | F5H1 | SALK_063792 | 3/2/05 | wild(.xls) mutant(.xls) |
4 , 87 | Yes | |
| At5g04330 | F5H2 | SALK_093419 | 3/2/08 | wild(.xls) mutant(.xls) |
4 , 90 | Yes | |
CCR & HCT Group |
|||||||
| Gene | Annotation | Accession | Date
Received |
IR
Spectra |
PCs, % Classified | Likely Spectrotype | Comments |
| At1g15950 | CCR1 | ||||||
| At1g80820 | CCR2 | ||||||
| At1g76470 | CCR-like1 | SALK_020615 | 3/2/05 | wild(.xls) mutant(.xls) |
5 , 68 | ||
| At2g02400 | CCR-like2 | ||||||
| At2g33590 | CCR-like3 | SALK_007759 | 3/2/05 | wild(.xls) mutant(.xls) |
4 , 65 | ||
| At2g33600 | CCR-like4 | ||||||
| At5g58490 | CCR-like5 | ||||||
| At5g48930 | HCT | ||||||
Back to top
Back to top
|
Back to top
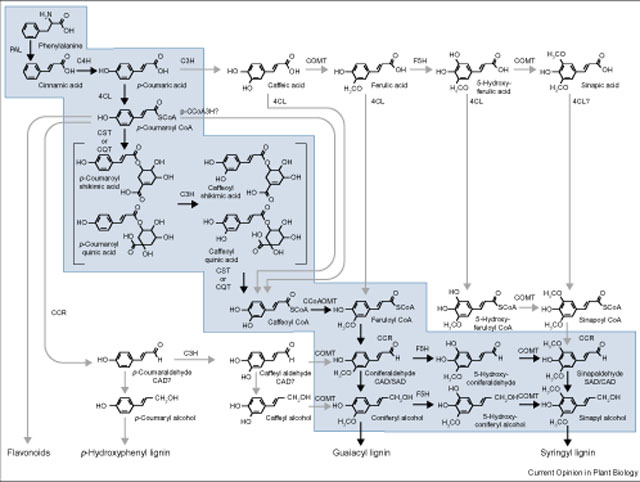
References:
Humphreys, J. M., C. C. S. Chapple. 2002. Rewriting the lignin roadmap. Plant Bio. 5, 224–229.
Back to top
Back to top
